  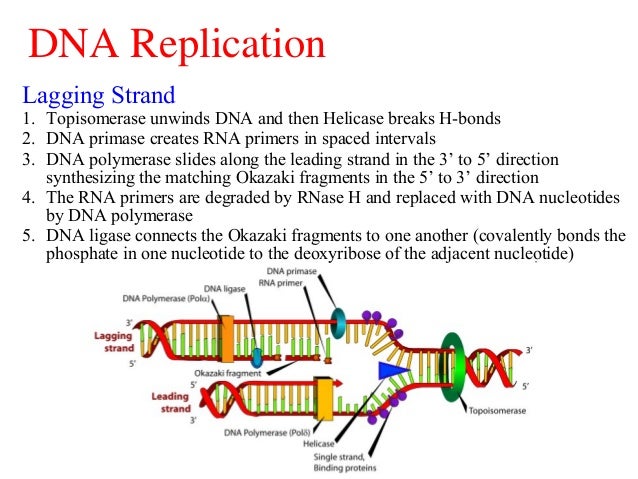
The analysis of Okazaki fragments offers a powerful approach to study not only DNA replication but also chromatin assembly pathways. In addition, global analysis of Okazaki fragments purified from asynchronous cultures reveals key properties that underlie genome-wide DNA replication ( 7). Consequently, Okazaki fragment ends correlate with nucleosome positions and have a periodicity reminiscent of the nucleosome repeat ( 6). Our published data shows that nucleosome assembly occurs very rapidly on nascent DNA and that polymerization by Pol δ is often impeded by nucleosomes. The second is a single-nucleotide resolution, deep sequencing approach that allows precise characterization of millions of individual DNA molecules ( 6 7). Using a doxycycline-repressible CDC9 allele, we have recently shown that unligated Okazaki fragments can be massively enriched ( 6) we have developed two assays to characterize them: The first is a low-resolution agarose gel based approach that allows us to visualize general properties of the population ( 6). 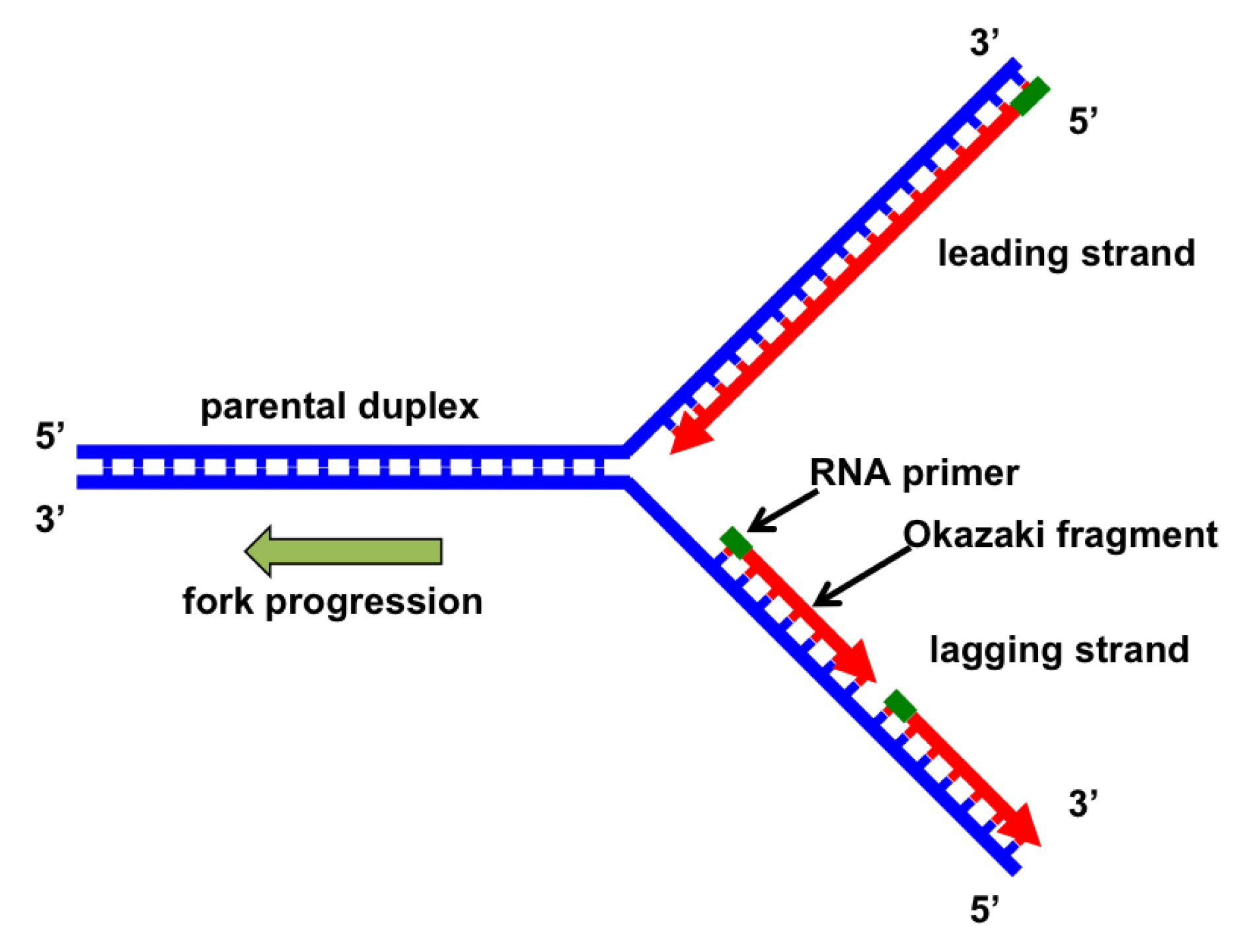
Strand displacement by Pol δ is stimulated by the structure specific Flap Endonuclease 1 (Fen1) nuclease, which cleaves the flap and generates a nick in the duplex DNA, which is ultimately sealed by the replicative ligase Lig1 (Cdc9)( 3- 5). Part of the RNA primer is likely removed by the action of RNase H, yet Pol δ also extends through the 5’ portion of the preceding fragment, displacing a single stranded flap( 3). Pol δ extends the 3’ end of the fragment until it meets the 5’ end of the preceding Okazaki fragment. After priming, elongation is stimulated by RFC (the clamp loader), which loads PCNA onto the 3’ primer end and exchanges Pol α-primase for the processive polymerase δ (Pol δ)( 2). Each Okazaki fragment is initiated by polymerase α-primase (Pol α-primase), which creates a short, ~35-50nt, mixed RNA-DNA primer( 1). Okazaki fragment synthesis on the lagging strand necessitates the repeated production of single stranded DNA and polymerization in the opposite direction to fork progression. However, conventional methods lack information about dynamics of individual replication forks and provide little insight into nucleosome organization during replication.ĭNA replication at the replisome is inherently asymmetric with the leading strand synthesized in advance of the lagging strand. Eukaryotic DNA replication has been traditionally studied using synchronization of cells followed by measurements of S-phase dependent incorporation of modified nucleotides, increases in copy number, single-stranded DNA or binding of specific replisome proteins. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
